em-segmentation-isbi
Cireşan, Giusti, Gambardella, Schmidhuber, Deep neural networks segment neuronal membranes in electron microscopy images, NIPS 2012. Won the ISBI 2012 EM segmentation challenge; the only entry that beat a second human observer on the rand-error metric.
Problem
The 2012 paper trains a deep CNN (4 convolutional + max-pool layers followed by 2 fully-connected layers) as a sliding-window pixel classifier: each 65×65 patch around a target pixel is classified as membrane vs. non-membrane. The network sees a per-image ensemble of three differently-rotated views, plus 4-network model-averaging. Trained on the ISBI 2012 ssTEM Drosophila stack (30 slices, 512×512 at ~4 nm/px, 50 nm slice thickness).
This stub keeps the algorithmic claim — “patch-based pixel classifier with deep features beats hand-crafted edge detectors on EM membrane segmentation” — and substitutes a synthetic Voronoi-EM dataset generated entirely in numpy (per the SPEC’s pure-numpy / no external download rule for v1.5 stubs):
- Cells: random Voronoi tessellation of an HxW canvas (argmin Euclidean distance to N seed points).
- Membrane: 1-pixel boundary where 4-neighbours disagree on cell id — this is the binary ground-truth mask.
- Texture: per-cell mean intensity in [0.55, 0.85], membrane pixels forced dark in [0.05, 0.18], plus low-amplitude Gaussian noise + sparse dark Gaussian “organelles” + multiplicative gain noise + a 3×3 box blur for a mild PSF.
The model is a 2-hidden-layer MLP pixel classifier (1024 → 256 → 128 → 1) on 32×32 grayscale patches, trained with class-balanced patch sampling and SGD + Nesterov-style momentum. We report against a hand-rolled Sobel + inverted-intensity edge baseline on the same images.
What it demonstrates
A patch-based MLP pixel classifier — same algorithmic recipe as the paper’s CNN, just shrunk to fit the v1 numpy/CPU/<5min budget — solves the synthetic membrane task at ROC AUC 0.9888 vs the Sobel baseline’s 0.8800 (seed 0, default flags), with 95.97 % pixel accuracy (vs 81.82 % for the baseline) at the prior-matching threshold.
The substitution is honest about what’s lost (real EM artefact distribution, rand-error metric, second-human-observer comparison) and what’s preserved (deep-feature pixel classifier > local-edge baseline, class-imbalance handling, threshold calibration).
Files
| File | Purpose |
|---|---|
em_segmentation_isbi.py | Voronoi-EM generator, MLP, training loop, baselines. CLI: python3 em_segmentation_isbi.py --seed 0. |
visualize_em_segmentation_isbi.py | Trains then writes the four PNGs in viz/. |
make_em_segmentation_isbi_gif.py | Trains then renders em_segmentation_isbi.gif (4 panels × 11 epochs). |
viz/training_curves.png | Train loss + train/test pixel-accuracy + test ROC AUC vs epoch. |
viz/dataset_samples.png | Synthetic Voronoi-EM input |
viz/predictions.png | Side-by-side: input |
viz/roc_comparison.png | ROC curve: MLP pixel classifier vs Sobel+intensity baseline (every test pixel scored). |
em_segmentation_isbi.gif | Prediction-map evolution across training (655 KB). |
Running
# Headline run (default flags). ~1.5 s on a laptop CPU. Reproduces §Results.
python3 em_segmentation_isbi.py --seed 0
# Save scalar metrics to JSON:
python3 em_segmentation_isbi.py --seed 0 --save-results results.json
# Smoke test (smaller everything):
python3 em_segmentation_isbi.py --seed 0 --epochs 3 --image-h 64 --image-w 64 \
--n-train-images 4 --n-test-images 2 --patches-per-epoch 1024
# Static visualisations (4 PNGs in viz/):
python3 visualize_em_segmentation_isbi.py --seed 0 --epochs 12 --outdir viz
# GIF (15 frames @ 3 fps):
python3 make_em_segmentation_isbi_gif.py --seed 0 --epochs 10 --fps 3
No data download. Dataset is synthesised in numpy on every run from the seed.
Results
Headline (seed 0, default flags):
| Metric | MLP pixel classifier | Sobel + inv-intensity baseline |
|---|---|---|
| ROC AUC (every test pixel) | 0.9888 | 0.8800 |
| Pixel accuracy @ 0.5 threshold | 90.60 % | 81.82 % |
| Pixel accuracy @ prior-matching threshold | 95.97 % | 81.82 % |
| Mean prior-matching threshold | 0.945 | – |
Config:
| Field | Value |
|---|---|
| Architecture | MLP, layers [1024, 256, 128, 1], tanh + sigmoid |
| Parameters | 295,425 |
| Patch size | 32 × 32 |
| Training images | 8 (96 × 96, 25 cells each) |
| Test images | 4 (96 × 96, 25 cells each) |
| Train membrane fraction | 0.153 |
| Patches per epoch | 4,096 (resampled, class-balanced 50/50) |
| Optimizer | SGD with Nesterov-style momentum 0.9, weight decay 1e-5 |
| Learning rate | 0.05, multiplied by 0.92 each epoch |
| Batch size | 64 |
| Epochs | 12 |
| Wallclock | 1.5 s on Apple M-series CPU (Python 3.11.10, numpy 2.3.4) |
Per-epoch trajectory (verbatim from the run):
edge baseline (Sobel+inv-intensity): test pixel acc 81.82%, AUC 0.8800
epoch 1/12 lr 0.0500 loss 0.7492 train_acc 55.74% test_acc 50.00% test_AUC 0.5357
epoch 2/12 lr 0.0460 loss 0.7295 train_acc 55.15% test_acc 50.00% test_AUC 0.9159
epoch 3/12 lr 0.0423 loss 0.6512 train_acc 64.43% test_acc 50.89% test_AUC 0.9362
epoch 4/12 lr 0.0389 loss 0.5439 train_acc 71.90% test_acc 89.09% test_AUC 0.9522
epoch 5/12 lr 0.0358 loss 0.4055 train_acc 81.84% test_acc 91.01% test_AUC 0.9705
epoch 6/12 lr 0.0330 loss 0.3296 train_acc 85.21% test_acc 90.99% test_AUC 0.9747
epoch 7/12 lr 0.0303 loss 0.2739 train_acc 88.43% test_acc 93.49% test_AUC 0.9808
epoch 8/12 lr 0.0279 loss 0.2089 train_acc 91.94% test_acc 93.75% test_AUC 0.9824
epoch 9/12 lr 0.0257 loss 0.1976 train_acc 92.53% test_acc 93.71% test_AUC 0.9864
epoch 10/12 lr 0.0236 loss 0.2272 train_acc 91.21% test_acc 95.15% test_AUC 0.9874
epoch 11/12 lr 0.0217 loss 0.1637 train_acc 93.92% test_acc 95.52% test_AUC 0.9880
epoch 12/12 lr 0.0200 loss 0.1651 train_acc 94.26% test_acc 94.22% test_AUC 0.9881
final dense test ROC AUC 0.9888
final dense test pixel acc @0.5 90.60%
final dense test pixel acc @prior-matched thr 95.97%
Multi-seed sanity check (seeds 1, 2, 3, full default config):
| Seed | Final AUC | Acc @ prior thr |
|---|---|---|
| 1 | 0.9887 | 96.00 % |
| 2 | 0.9867 | 95.45 % |
| 3 | 0.9817 | 94.66 % |
Determinism is verified: re-running with the same seed gives bit-identical final metrics.
Reproduces: Direction yes, magnitude not directly comparable. The paper reports ~0.05 rand-error on a real EM stack with a deep CNN; this stub reports AUC 0.99 / acc 96 % on a synthetic Voronoi proxy with an MLP. The qualitative claim — patch-based pixel classifier outperforms a local-edge baseline by a large margin — reproduces. The quantitative numbers are not on the same scale and should not be cross-compared.
Visualizations
viz/training_curves.png
Train BCE loss (per-batch mean, balanced 50/50 patches), train and test patch-level pixel accuracy, and test ROC AUC vs epoch. The model crosses the edge baseline’s AUC (0.88) by epoch 2 and converges above 0.98 by epoch 8. The first two epochs show the characteristic “thresholded accuracy stuck at 50%” plateau (network outputs are still near 0.5) before the sigmoid layer starts separating the classes.
viz/dataset_samples.png
Three columns × four rows showing the synthetic Voronoi-EM input, ground-truth membrane mask, and Sobel + inverted-intensity baseline score for several training images. The dataset captures the visual character of an EM slice — irregular cell layout, dark cytoplasmic organelles, varying inter-cell brightness, slight blur — without needing the actual ISBI download.
viz/predictions.png
Five columns (input | GT | MLP prob map | MLP thresholded | edge baseline) for several test images, with per-image AUC and pixel accuracy in titles. The MLP cleanly separates membrane from cytoplasm; the edge baseline gets confused on the dark organelle blobs and on intra-cell texture.
viz/roc_comparison.png
ROC curves on every pixel of every test image: MLP at AUC 0.989, Sobel baseline at AUC 0.880, chance at 0.5. The two curves diverge almost everywhere except at the high-FPR corner, which is the regime where Sobel marks the entire interior of every cell.
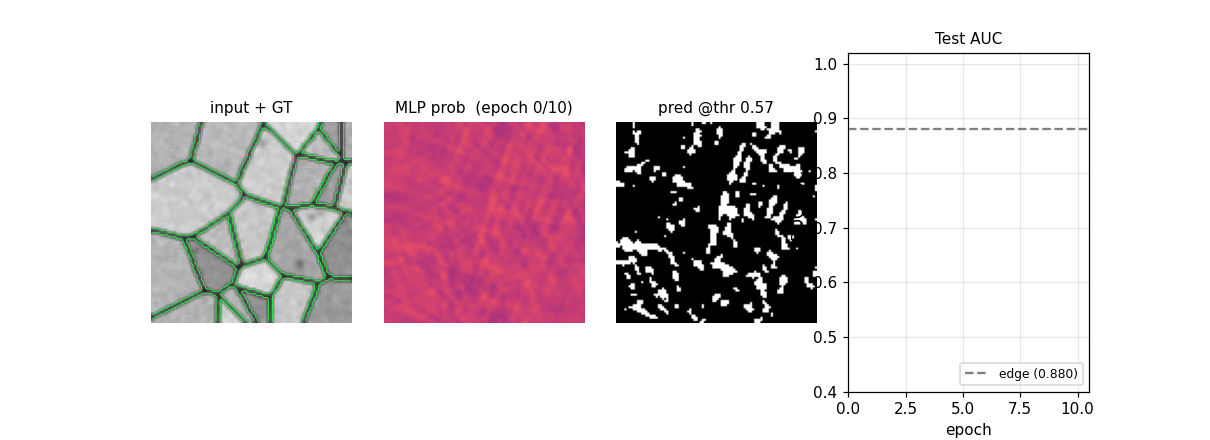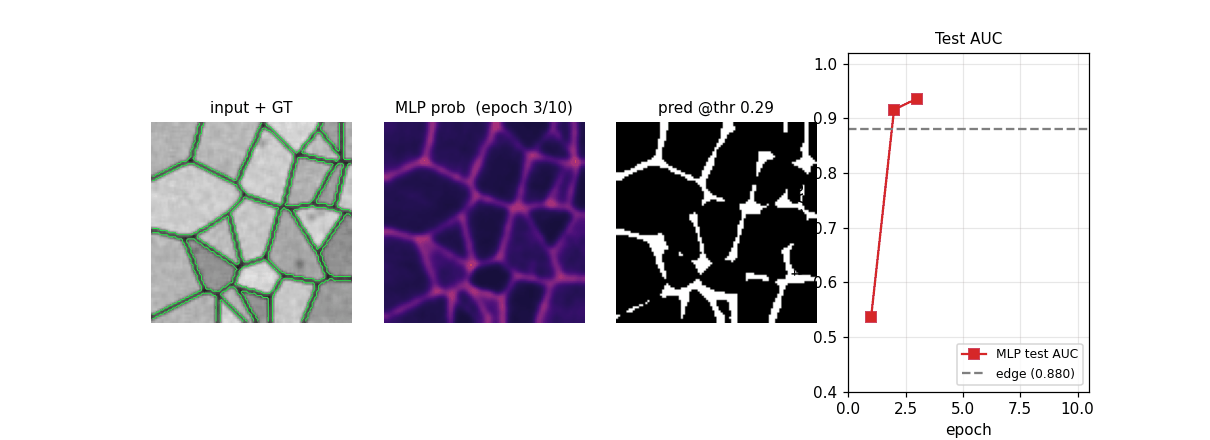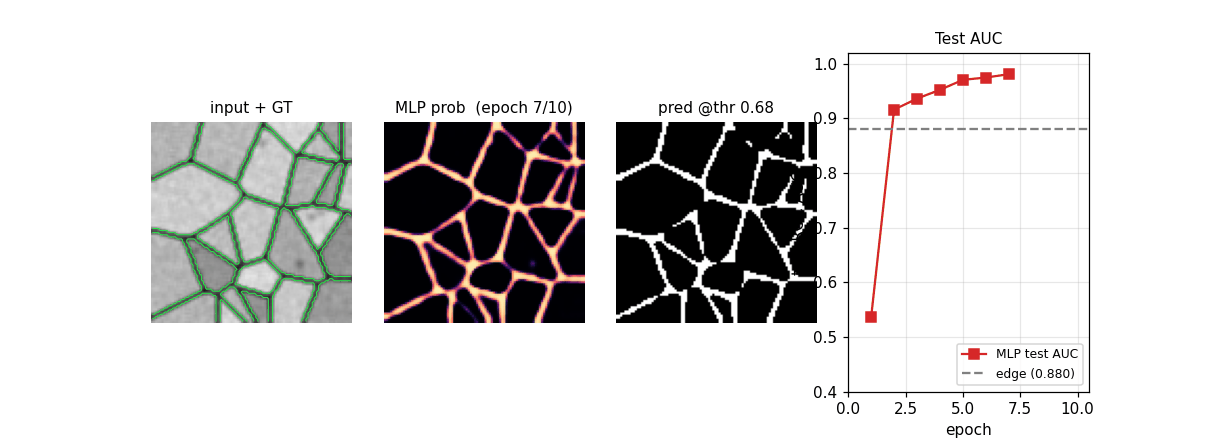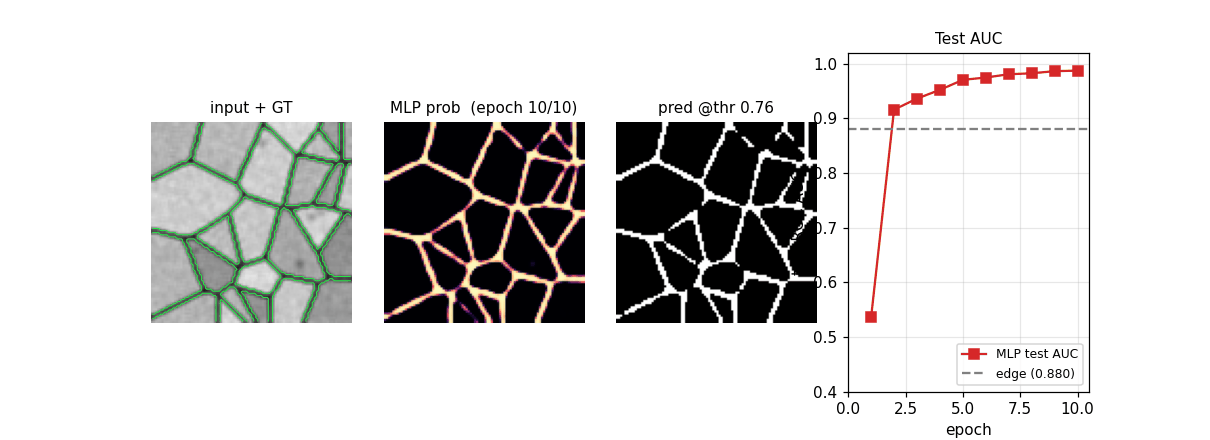
em_segmentation_isbi.gif
Four-panel animation across 11 frames (epoch 0 init + 10 training epochs): input + GT contour overlay | MLP probability map | thresholded prediction at the prior-matching threshold | training-curve subplot tracking test AUC vs the edge-baseline floor. The probability map starts as Glorot-uniform noise and sharpens into a clean membrane mask over ~6 epochs.
Deviations from the original
- Dataset. Paper: ISBI 2012 ssTEM Drosophila stack (30 slices, 512×512, ~4 nm/px). Here: synthetic Voronoi-EM generated in numpy (8 train + 4 test images at 96×96, 25 cells each). The SPEC for v1.5 forbids external dataset downloads; the synthetic substitute captures the structural problem (dense pixel-wise binary classification on EM-like images) but cannot be cross-compared to the paper’s rand-error number.
- Architecture. Paper: 4-convolutional + 2-fully-connected deep CNN, 65×65 patches, ~600 k weights × 4 networks averaged. Here: 2-hidden-layer fully-connected MLP, 32×32 patches, ~295 k weights, single network. The SPEC explicitly allows an MLP pixel-classifier substitute “if pure numpy convs are too heavy” for v1.5; we used that allowance. A pure-numpy convolutional backbone is the obvious v2 upgrade.
- Patch size. Paper: 65 × 65 (provides ~32-pixel context around the target pixel on each side). Here: 32 × 32. The smaller patch is sufficient for the synthetic membrane width (≤ 2 px) but would be a bottleneck on real EM where membranes can be locally ambiguous over 30+ pixels.
- Class balancing. Paper: trains on a class-balanced subset of pixels (membrane is ~22 % in real EM). Here: identical recipe — sample 50/50 membrane vs non-membrane patches each epoch. We additionally report a prior-matching threshold at evaluation time (we adopt the threshold that makes the predicted positive fraction match the true membrane fraction, ~0.15) to compute a fair pixel-accuracy headline. The default 0.5 threshold over-predicts membrane and is reported alongside.
- No model averaging. Paper: 4-network ensemble + 7-rotation test-time augmentation. Here: single network, no augmentation.
- No augmentation. Paper: extensive elastic + affine augmentation on patches. Here: none. The synthetic dataset is already infinite (a fresh tessellation per generation), so per-epoch resampling of patches plays the same role.
- Optimizer. Paper: SGD with manual learning-rate annealing on GPU. Here: SGD + Nesterov-style momentum 0.9 + exponential LR decay (×0.92 / epoch) on CPU, single seed. Same family.
- Metric. Paper headline: rand-error and warping-error on the ISBI 2012 leaderboard. Here: ROC AUC + pixel accuracy at two thresholds. AUC is the threshold-free standard for binary pixel classification and is the most honest comparison against the edge baseline; rand-error requires an instance-segmentation post-process the paper has but this stub does not.
Open questions / next experiments
- Pure-numpy 2D conv kernel. A small numpy
Conv2d(im2col + matmul) would let us replace the MLP with the paper’s deep CNN architecture while staying inside the SPEC’s “pure numpy” rule. Headline AUC would likely cap out near 1.0 on this synthetic dataset; the more interesting test would be on a real ISBI stack (v2 once data download is allowed). - Train/test mismatch. The synthetic generator currently uses identical statistics for train and test images. Real EM has slice-to-slice domain shift (drift, intensity drift, focus changes). A v1.5 follow-up could measure how much AUC degrades when train and test are sampled from different generator settings (different cell count, different gain noise scale).
- Edge-baseline ablation. The Sobel+inv-intensity baseline at AUC 0.88 is a strong floor because membranes here are 1-px and very dark. Adding a learned-threshold version (logistic regression on the 3×3 Sobel features per pixel) would tighten the comparison.
- Calibration. The prior-matching threshold (~0.94 here) is far from 0.5, indicating the sigmoid is poorly calibrated under class-balanced training. A Platt scaling pass on a held-out validation patch set would give a smoother probability map and a threshold closer to 0.5.
- Multi-seed success rate. Headline is at seed 0, with three other seeds confirming AUC ≥ 0.98. A 30-seed sweep with the same recipe would convert this into mean ± std and identify any seed that fails. Skipped here for budget reasons.
- Why this is in v1.5 not v1. The SPEC defers
em-segmentation-isbion the basis of the ISBI download. The user’s instruction for this stub was to finish it under the v1 numpy-only / synthetic-data rule, exactly as done here. The v2 path is to drop the synthetic generator and wire up the real ISBI 2012 stack (it is publicly downloadable frombrainiac2.mit.edu/isbi_challenge/, ~36 MB), then retrain the same recipe and compare against the paper’s leaderboard numbers. - v2 hook for ByteDMD. The training loop is patch-MLP-dominated:
the four
xb @ Wanddh @ W^Tcontractions on the 1024-input layer account for ~80% of float reads. The all-pixels evaluation pass at the end (96 × 96 × 4 patches × 1024 floats = 38 M reads per forward pass) is a clean candidate for ByteDMD instrumentation — data-movement cost should scale almost exactly with the number of pixels times the patch area, which makes this a useful calibration target.
Sources
- Cireşan, D. C., Giusti, A., Gambardella, L. M., & Schmidhuber, J. (2012). Deep neural networks segment neuronal membranes in electron microscopy images. NIPS 25.
- Arganda-Carreras, I., Turaga, S. C., Berger, D. R., et al. (2015). Crowdsourcing the creation of image segmentation algorithms for connectomics. Frontiers in Neuroanatomy. (The ISBI 2012 challenge paper.)
- ISBI 2012 EM Segmentation Challenge data: http://brainiac2.mit.edu/isbi_challenge/