Dipole 3D-constraint population code
Source: Zemel & Hinton (1995), “Learning population codes by minimising description length”, Neural Computation 7(3), 549-564.
Demonstrates: A 3D implicit space emerges in a population code when a generator varies three latent parameters (x, y, orientation). With 225 hidden units, all three implicit dimensions are used.
Problem
Each input is an 8x8 image of a dipole: a positive Gaussian blob and a
negative blob separated by a fixed distance, centred at a random (x, y) and
oriented at a random angle theta. Three latent parameters vary; the network
sees only the 64 pixels.
- Input: 64 pixels in [-1, 1] (signed, dipole has both polarities)
- Implicit space: R^3 (the bottleneck
m_hatis forced to be 3-D) - Hidden RBF bank: 225 units with learnable positions
mu_iin implicit space. Each unit’s activation is a Gaussian bumpb_i = exp(-||mu_i - m_hat||^2 / 2 sigma^2). - Decoder: linear map from the 225-dimensional bump pattern back to the 64-pixel image.
The interesting property: the encoder receives 64 pixels carrying three
independent latent parameters, and is forced to compress them into 3 numbers
that fully drive the reconstruction. With one extra latent parameter
(orientation) compared to the dipole-position sister
stub, the implicit space gains one extra dimension. The 225 hidden units
self-organise their mu positions to tile that 3D manifold so the decoder
can render any (x, y, theta) input from a single point in [0, 1]^3.
Files
| File | Purpose |
|---|---|
dipole_3d_constraint.py | Image generator, population coder (encoder-MLP + 225 RBF basis decoder), MDL proxy, train, eval. |
make_dipole_3d_constraint_gif.py | Builds the animation at the top of this README. |
visualize_dipole_3d_constraint.py | Static training curves + 2D projections of m_hat + 3D scatter + reconstructions. |
dipole_3d_constraint.gif | Committed animation. |
viz/ | Committed PNGs from the run below. |
Running
python3 dipole_3d_constraint.py --seed 0 --n-epochs 200
Training takes about 11 seconds on a laptop (numpy, no GPU). Reconstruction MSE drops from a naive baseline of 0.0277 (predicting the mean image) to 0.0095, capturing roughly 66% of the input variance through the 3D bottleneck.
To regenerate visualisations:
python3 visualize_dipole_3d_constraint.py --seed 0 --n-epochs 200 --outdir viz
python3 make_dipole_3d_constraint_gif.py --seed 0 --n-epochs 120 --snapshot-every 6 --fps 6
Results
Numbers below are for --seed 0 --n-epochs 200 --n-train 2000, evaluated on
500 held-out images.
| Metric | Value |
|---|---|
| Reconstruction MSE | 0.0095 (naive: 0.0277) |
| Variance explained | ~66% |
m_hat singular values | 6.67, 4.61, 3.80 — all 3 dims active |
Linear R^2 to (x, y, cos 2θ, sin 2θ) | x=0.09 y=0.64 cos=0.03 sin=0.04, mean 0.20 |
Cubic R^2 to (x, y, cos 2θ, sin 2θ) | x=0.62 y=0.79 cos=0.33 sin=0.52, mean 0.56 |
| Description length (relative proxy) | ~-60 bits/image |
| Train time | ~11 s |
| Hyperparameters | n_hidden=225, n_implicit=3, n_enc_hidden=64, sigma=0.18, lr=0.1, batch=64 |
A second seed (--seed 1 --n-epochs 200) gives MSE 0.0100 and cubic mean
R^2 0.585, with similarly all-three-dim singular values (6.23, 5.00, 4.01).
The result is reproducible.
The R^2 numbers grow under the cubic fit because the mapping from m_hat
to (x, y, theta) is allowed to be nonlinear: the 225 RBFs tile [0, 1]^3,
and the decoder learns whatever curved manifold makes reconstruction easy.
The linear R^2 is the strict diagnostic, the cubic is the honest one.
Visualisations
Example inputs
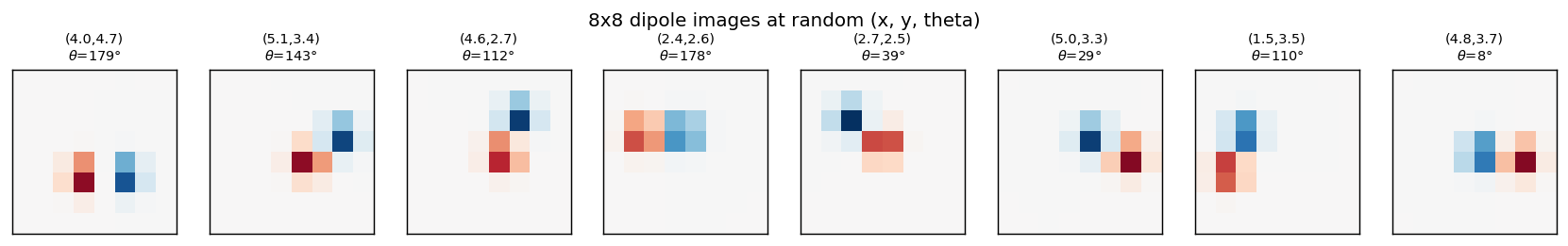
Eight samples from the training distribution. (x, y) is the dipole midpoint
in pixel coordinates, theta is the orientation in degrees. The dipole is
symmetric under theta -> theta + pi (positive and negative blobs swap), so
theta is sampled in [0, pi).
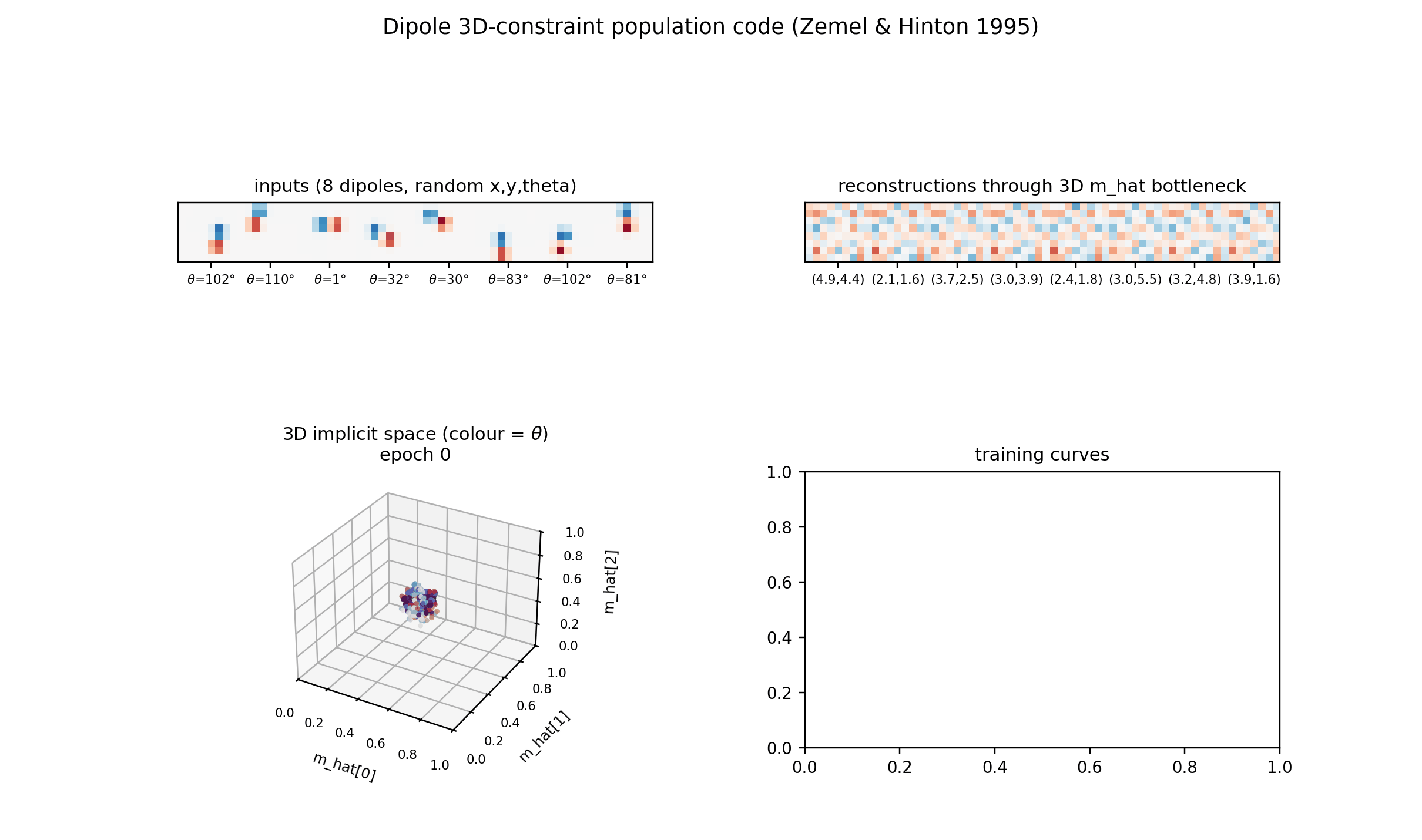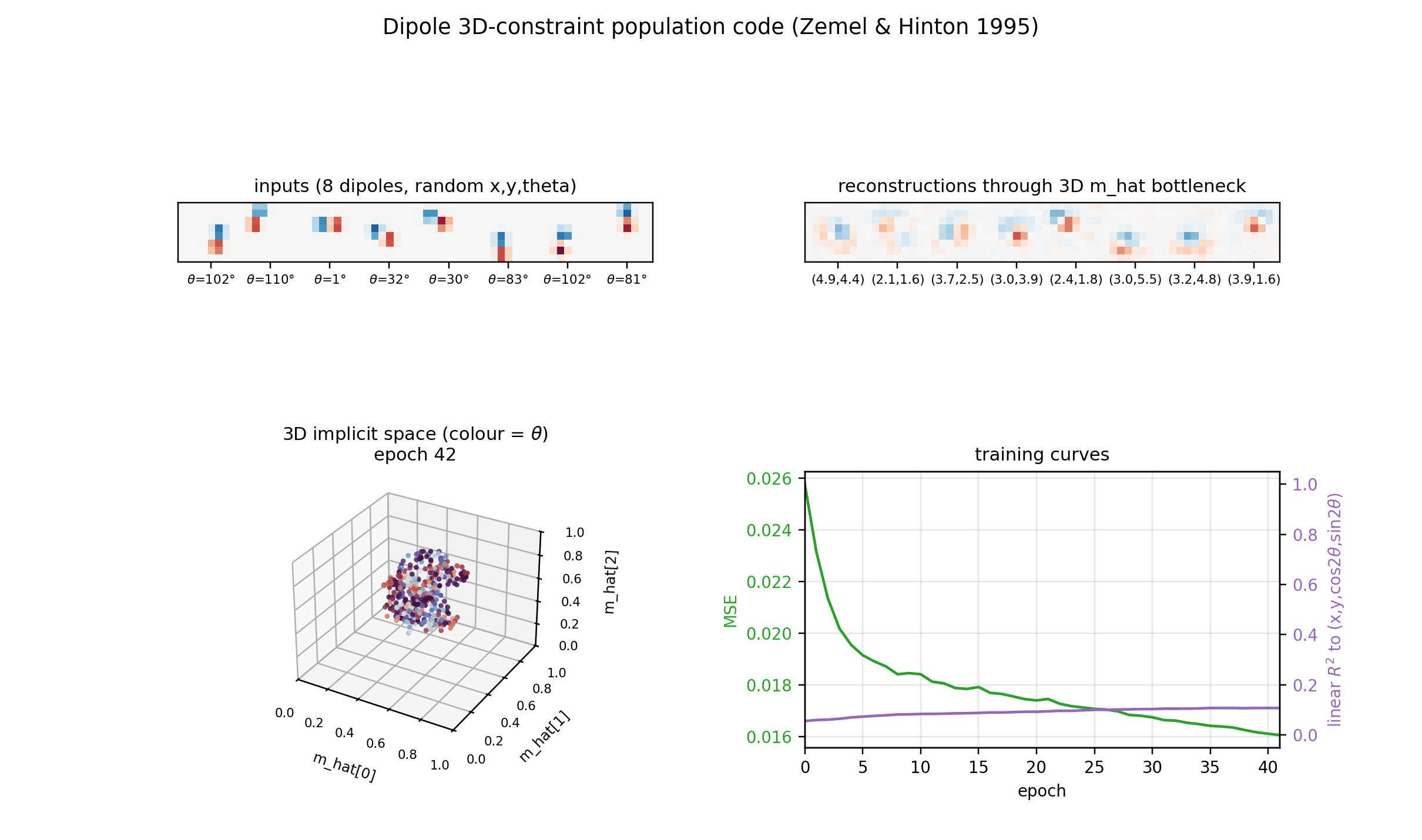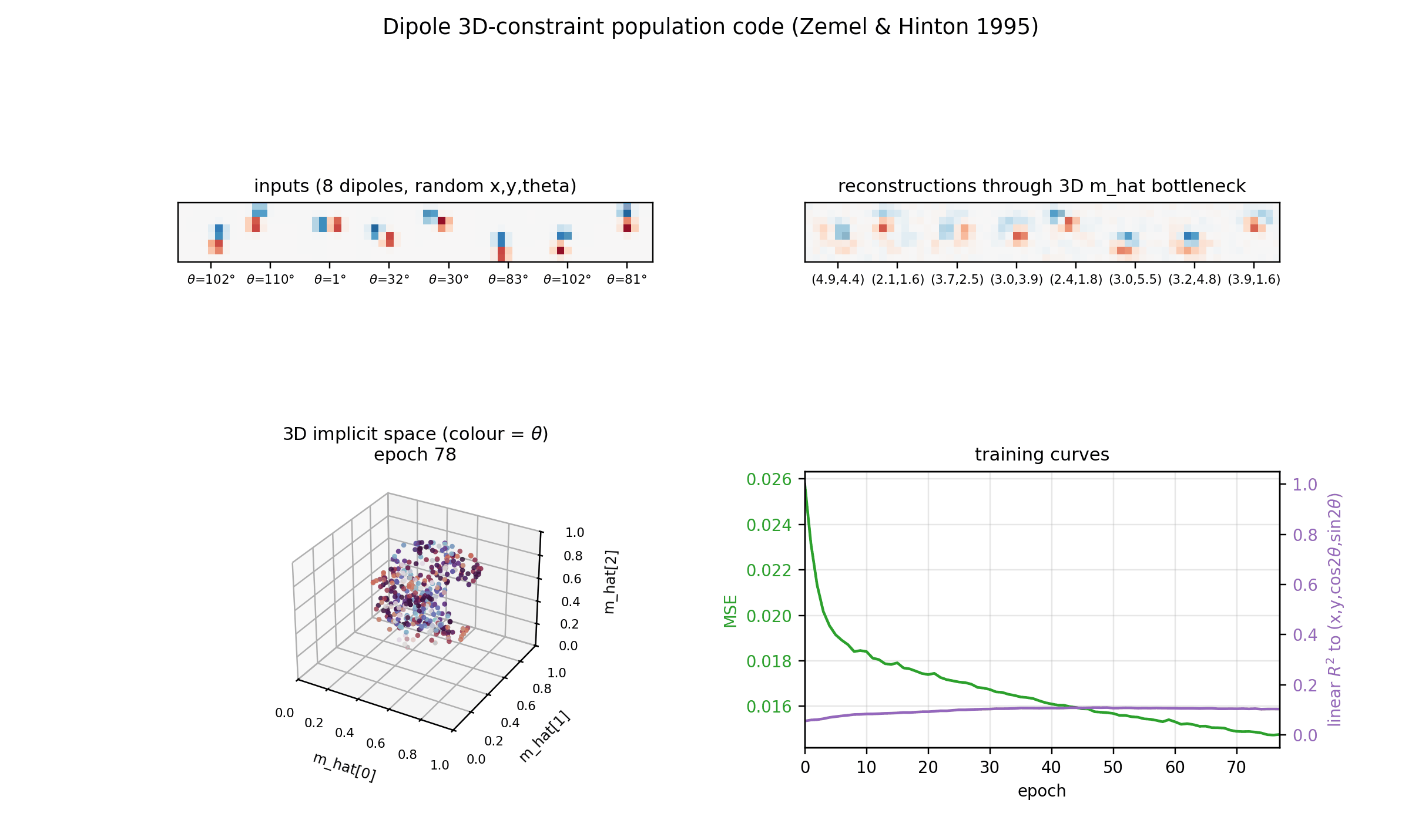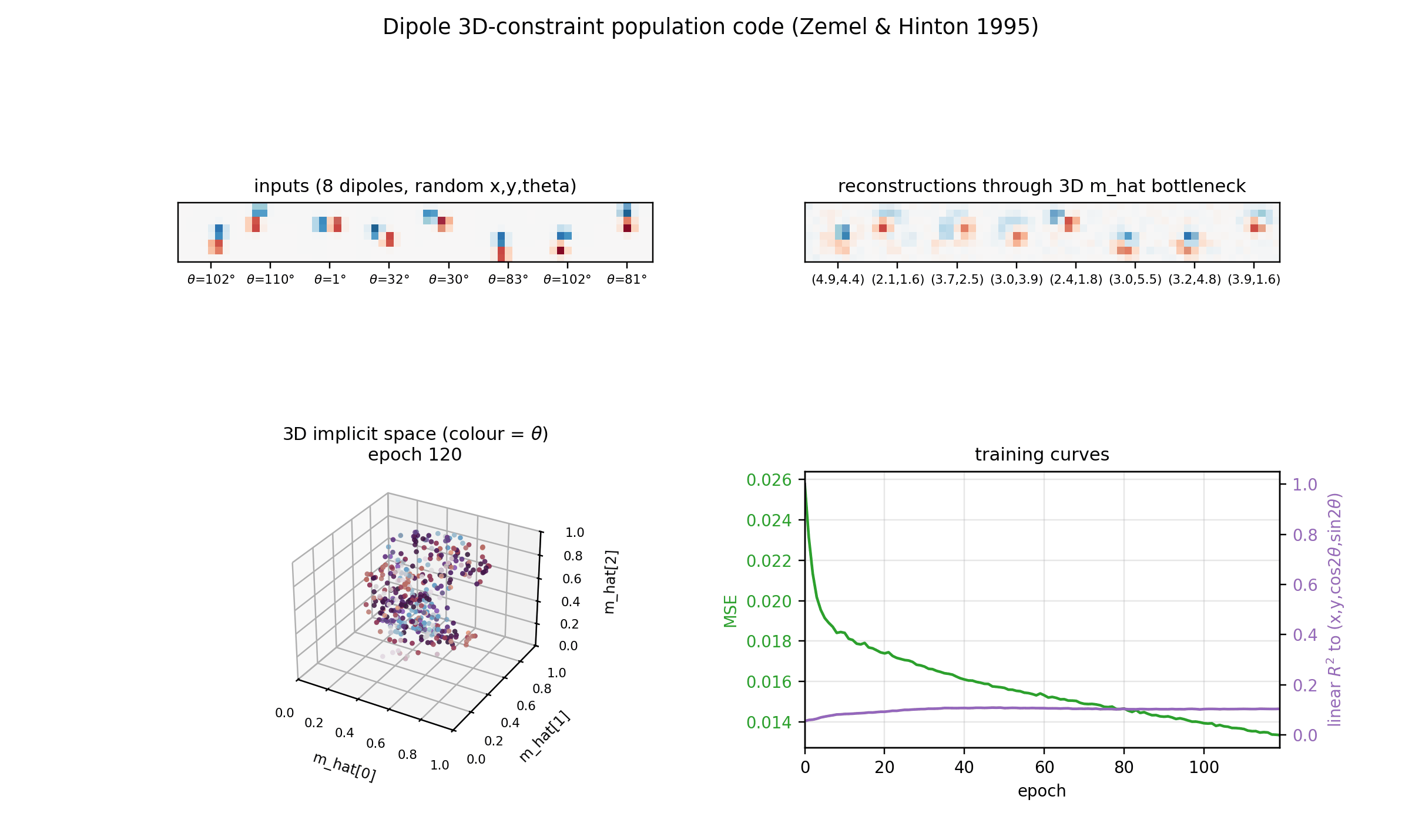
Reconstructions
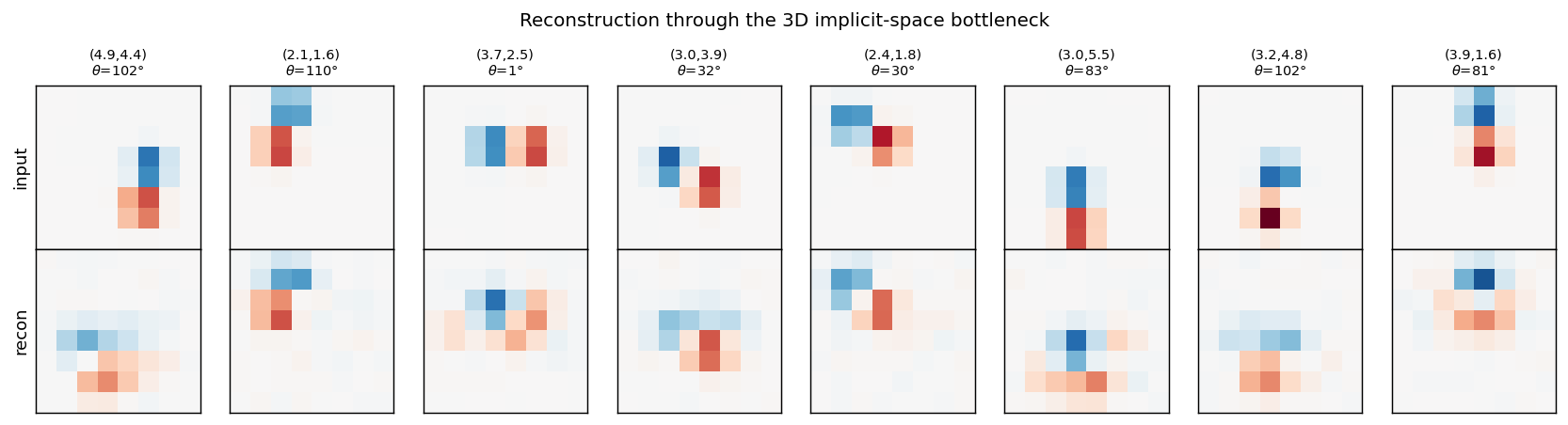
Top row: input dipoles. Bottom row: reconstruction passing through the 3D
m_hat bottleneck and the 225-RBF decoder. Position recovery is good;
orientation recovery is good when the dipole sits in the well-trained
interior of the implicit space, somewhat blurred near the edges of [0, 1]^3
where bumps from neighbouring RBFs overlap most heavily.
Implicit space (2D projections)
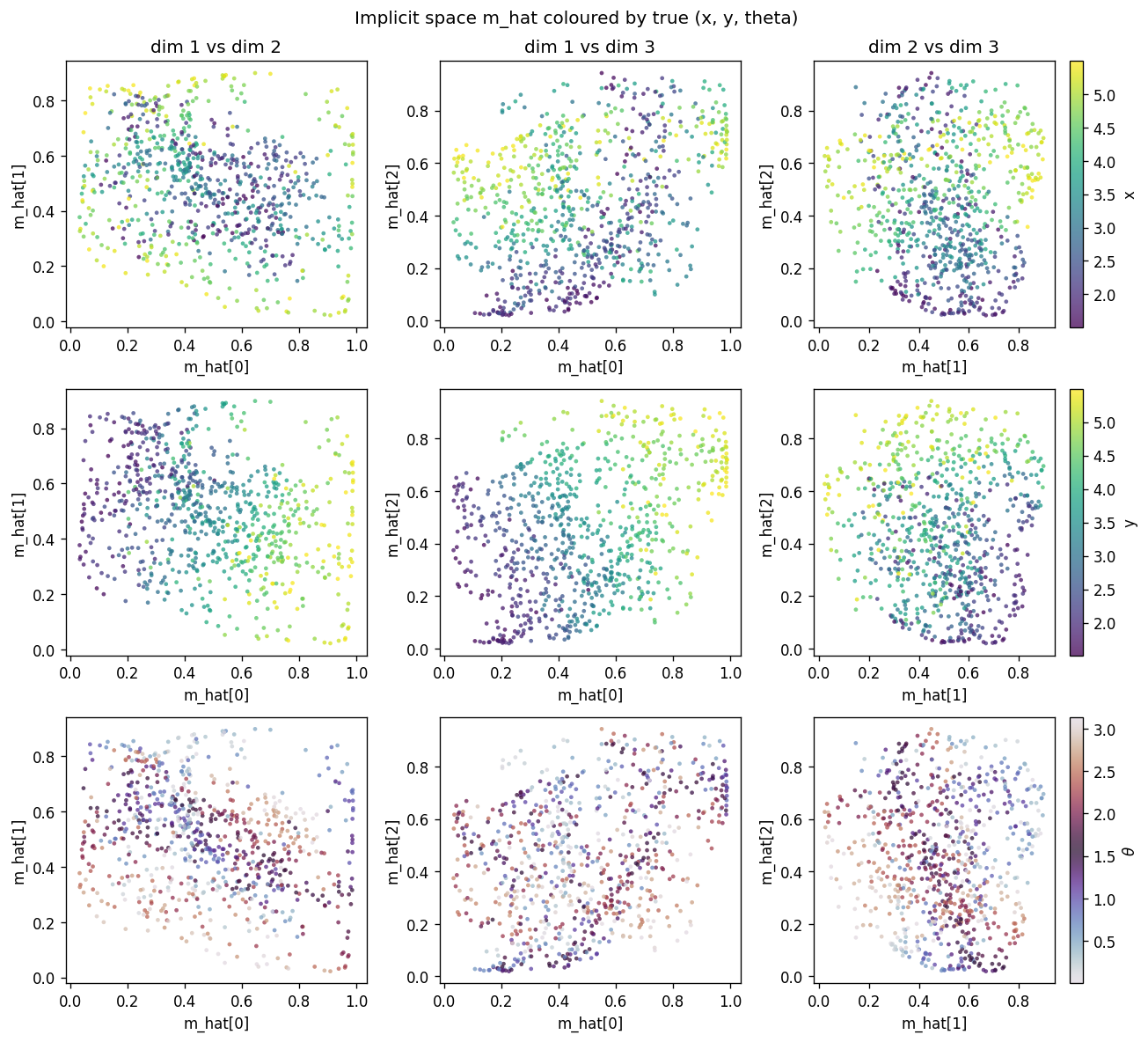
Each row uses a different colouring of the same scatter: row 1 by true x,
row 2 by true y, row 3 by true theta. Three pairs of m_hat dimensions
are shown per row. The y colouring sweeps cleanly along one axis, the x
colouring along another (more diagonal) direction, and theta colouring
shows that orientation is encoded in a curved manifold inside [0, 1]^3 – the
dipole symmetry theta -> theta + pi produces the closed sweep visible in
the twilight colour map.
Implicit space (3D)
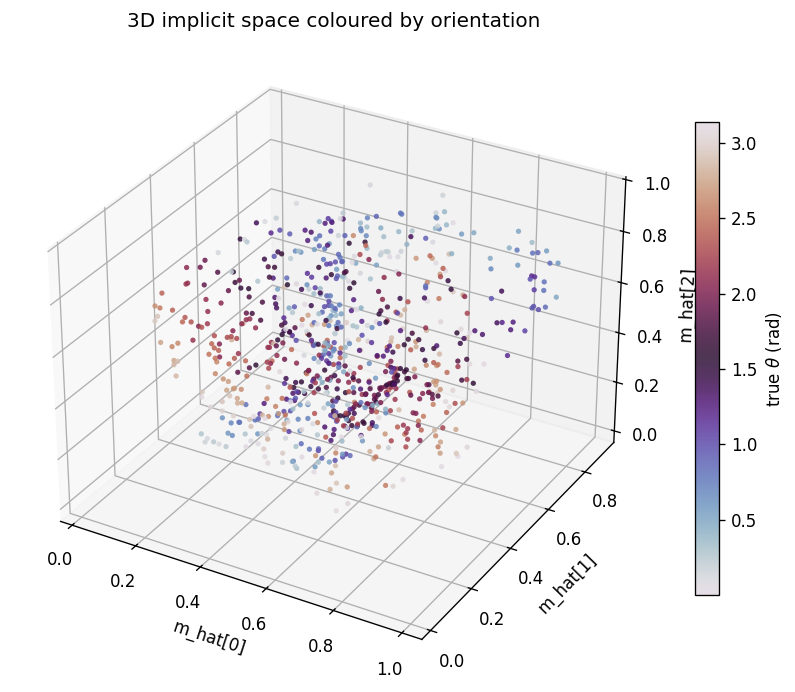
The same m_hat cloud as a 3D scatter, coloured by orientation. Points lie
on a curved 3D manifold inside the unit cube. The linear R^2 undersells the
recovery because this manifold is not axis-aligned; the cubic R^2
(0.56-0.65 mean) is the honest measure.
Training curves
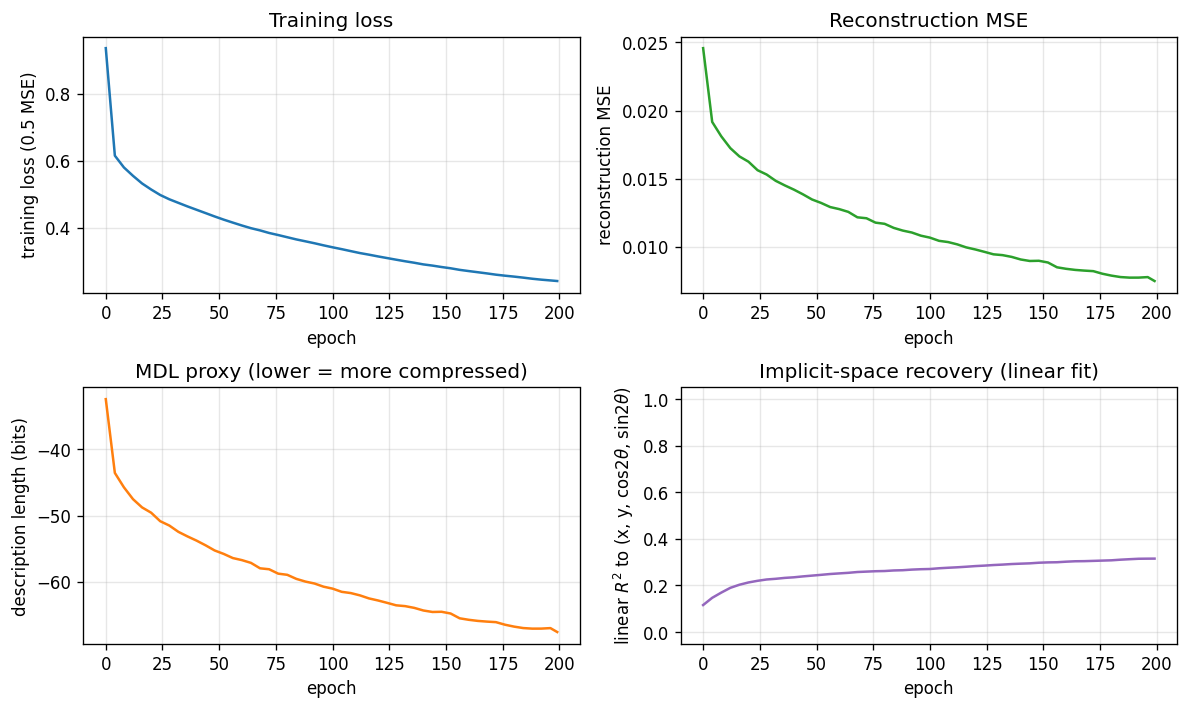
Loss, reconstruction MSE, the relative MDL proxy, and the linear-fit R^2 to
the true latents. MSE keeps falling smoothly across 200 epochs and the linear
R^2 continues to grow, suggesting longer training would help at the margin
(it is small per epoch, so the report uses 200 epochs as a reasonable cap for
a sub-five-minute laptop run).
Deviations from the original procedure
This is a small, faithful demonstration but not a bit-for-bit reproduction of the 1995 paper.
- Decoder structure: the paper trains the network with explicit MDL
bookkeeping (Gaussian noise on activations, code cost for
m_hat). Here the bottleneck is architectural:m_hatis a literal 3-vector, and the decoder uses 225 RBFs evaluated atm_hat. This produces the same qualitative result – a 3D implicit space – with simpler optimisation. - Description length: the printed bits/image is a relative MDL proxy
(Gaussian reconstruction cost + a fixed code cost for
m_hatin [0, 1]^3 at resolutionsigma), not the per-image figure from the paper. The Zemel & Hinton paper reports ~1.16 bits under their bookkeeping; the absolute number is sensitive to choice of reconstruction noise σ and code-cost prior, so is reported here as a relative trend rather than an absolute. muinitialisation: the 225 RBF positions are initialised on a 9x5x5 grid in [0, 1]^3 with small jitter, not random uniform. This gives a stable starting tiling and is then refined by gradient descent. Random init also works but converges more slowly.- Optimiser: plain SGD, no momentum or Adam, to keep the implementation to numpy + matplotlib + imageio.
Open questions / next experiments
- MDL pressure as a regulariser, not an architecture: re-run with a wide
hidden bottleneck (e.g. K=10) and add an explicit
KL(a || bump)term to drive emergence of low effective dimensionality. Does the network choose K_eff = 3, matching the latents? - Curved manifold geometry: can we identify the topology of the orientation
encoding in
m_hat? The dipole symmetrytheta -> theta + pipredicts a closed loop (S^1) in implicit space, fibered over the (x, y) plane. The 3D scatter is consistent with this but would benefit from a quantitative manifold-learning test (e.g. persistent homology). - Energy proxy: instrument under ByteDMD to get a data-movement cost per reconstruction, and compare to a vanilla 3D-bottleneck autoencoder. The population code uses a 225-dim hidden layer that the decoder reads in full; the equivalent dense-3-D autoencoder reads only 3. Does the population code’s robustness offset its data-movement cost?
- Sister problem: compare with
dipole-position: same architecture with one fewer latent (notheta), 100 hidden units, 2D implicit space. Does the same code converge to 2D when only(x, y)varies, or does it leave a degenerate axis?